Different types of chromatin PowerPoint PPT Presentation
1 / 51
Title: Different types of chromatin
1
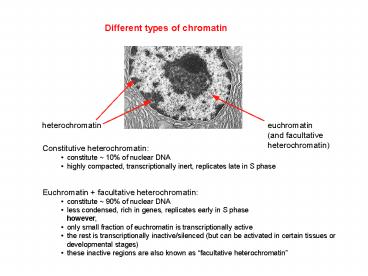
Different types of chromatin
- Constitutive heterochromatin
- constitute 10 of nuclear DNA
- highly compacted, transcriptionally inert,
replicates late in S phase
- Euchromatin facultative heterochromatin
- constitute 90 of nuclear DNA
- less condensed, rich in genes, replicates early
in S phase - however,
- only small fraction of euchromatin is
transcriptionally active - the rest is transcriptionally inactive/silenced
(but can be activated in certain tissues or - developmental stages)
- these inactive regions are also known as
facultative heterochromatin
2
Gene silencing and why is it important
- In any given cell, only a small percentage of
all genes are expressed - vast majority of the genome has to be shut down
or silenced - knowing which genes to keep on and which ones
to silence is - critical for a cell to survive and proliferate
normally
3
Gene silencing and why is it important
Wolffe and Matzke, Science, 1999
4
Epigenetics and development
n
n
2n DNA content
Differentiation
same DNA content, gt 200 cell types
5
Epigenetics and development
2n DNA content
De-differentiation?
same DNA content, gt 200 cell types
examples
- Cloning by nuclear transfer --gt regenerate entire
organism from transfer of single nucleus (e.g.
Dolly) - Induced pluripotent stem cells (iPS) --gt
expression of 4 genes are sufficient to transform
differentiated cells to stem cells - Both processes must involve reprogramming of
epigenome!
6
Epigenetics and epigenetic regulation
Definition of Epigenetics
- heritable changes in gene expression that do
not involve changes - in DNA sequences
- mechanisms
- DNA methylation
- histone modifications
- examples
- Developmentally regulated or tissue specific
gene expression - X chromosome dosage compensation
- Drosophila position effect variegation (PEV)
7
Epigenetic mechanism 1 DNA methylation
- DNA methylation has long been correlated with
repression of gene expression - DNA methylation mostly occurs on CpG
dinucleotides
DNMTs
methyl group is added to the cytosine
methylation status is maintained during
replication by DNMTs
8
DNA methylation and gene silencing
9
A class of proteins called MBD bind methylated DNA
- MeCP2 is the first protein found to bind to
methylated DNA - mutation of MeCP2 gene causes Rett Syndrome in
humans
shifted probes
unmethylated probe
methylated probe
10
MBD proteins interact with histone deacetylases
- MBD2 and HDACs co-purify
- in the same complex
- MBD2 physically co-IPs
- with HDACs
- MBD2 co-IPs with
- HDAC activity
11
DNA methylation recruits histone deacetylases
12
Epigenetic mechanism 2 histone methylation
- histone H3 is methylated at several lysine
residues - H3 K4-methylation is associated with
transcriptional activation - whereas K9-, K27-methylation is associated
with repression - these H3 methylation sites define the
transcriptional/epigenetic states - of the associated genes/chromatin domains
13
Epigenetics example 1 Tissue-specific and
developmentally regulated gene expression
- globin genes are expressed only in erythroid
cells - hemoglobin made up of 2 copies each of a- and
b-chains
14
Gene order of globin clusters mirror expression
pattern during development
15
Globin genes are tissue-specific and
developmentally regulated
- Distinct isoforms of the globin genes are
expressed at different developmental stages - e.g., for the b-globin family, expression goes
from e- to g- to b-isoforms
- mutations in adult isoforms of globin genes
result in thalassemia
16
Globin LCR and adult b-globin promoters are
hyperacetylated in adult mouse erythroid
leukemia cells upon induction
Forsberg et al, PNAS, 2000
17
Epigenetics example 2 Dosage compensation of X
chromosome
- for many organisms, females have 2 copies of
the X chromosome whereas males - only have single copy
- how to balance expression dosage of X-linked
genes?
18
Drosophila polytene chromosomes
- Drosophila genome has 4 chromosomes
- polytene chromosomes result from
endoreplication - (DNA replication without cytokinesis)
- giant chromosomes that are easily visible
2048 identical DNA strands
19
X chromosome in Drosophila
- the X chromosome of male Drosophila is
transcriptionally twice as active - increased transcription of the active X
chromosome is marked by - hyper-acetylated histones
20
X chromosome inactivation
- In female mammals, one of the two X chromosomes
in the genome is transcriptionally inactivated - in order to equalize expression of X-linked
genes in males and females (dosage compensation) - Inactivation of the maternal or paternal
chromosome is random
21
X chromosome inactivation
- In X inactivation, almost the entire X
chromosome is transcriptionally silenced - Transcriptional silencing of this chromosome
correlates with distinct histone modification
patterns - eg. histone H4 is hypo-acetylated on the
inactive X chromosome
metaphase chromosome immunofluorescence
Jeppesen et al, Cell, 1993
22
(No Transcript)
23
X inactivation involves sequential epigenetic
modifications of the silenced chromosome
24
Epigenetics example 3 Position effect
variegation in Drosophila
White gene encodes red pigment in eye
25
Position effect variegation in Drosophila
example of epigenetic regulation since silencing
of white gene is NOT due to DNA mutation, but
due to translocation and spreading of
heterochromatin
26
Position effect variegation in Drosophila
Su(var) mutations Suppressors of PEV e.g.
Su(var)2-5 HP1 Su(var)3-9 SET-domain
protein
27
Identification of H3 Lys9 methyltransferase
- The first lysine-specific HMT was identified by
IP-in vitro activity assays
1
9
- The SET domain of the SUV39H1 is required for
histone methyltransferase activity - and this enzyme methylates H3 at Lys9
Rea et al, Nature, 2000
28
Identification of other H3 methyltransferases
- The SET domain is the conserved catalytic core
of histone methyltransferases
...
...
ARTKQTARKSTGGKAPRK
ARKSA
H3
9
4
27
Me
Me
Me
Suv39H1/2 Su(var) 3-9
human
Drosophila
SET domain
29
Identification of H3 methyltransferases
- The SET domain is the conserved catalytic core
of histone methyltransferases
...
...
ARTKQTARKSTGGKAPRK
ARKSA
H3
9
4
27
Me
Me
Me
MLL Trx
Suv39H1/2 Su(var) 3-9
EZH2 E(Z)
human
Drosophila
SET domain
30
How does H3 K9-methylation functions in
heterochromatin assembly?
- back to early genetics studies in Drosophila
- Su(Var) 2-5 (gene) codes for heterochromatin
protein 1 (HP1)
- HP1 in Drosophila is localized to the
chromocenter
HP1
DNA
31
Ectopic expression of SUV39H1 causes
redistribution of HP1
Melcher et al, MCB, 2000
32
Lys9-methylated H3 binds to the conserved motif
called chromodomain
- Using the peptide pull-down assay, it was found
that Lys9-methylated H3 binds to - heterochromatin protein 1 (HP1)
- HP1 is a protein previously identified to be
enriched in and important for - heterochromatin assembly
- Lys9-methylated H3 binds to HP1 via the
chromodomain motif in HP1
Bannister et al, Nature, 2001
33
H3 K9-methylation is required for HP1 localization
Lachner et al, Nature, 2001
34
H3 K9-methylation is required for HP1 localization
Lachner et al, Nature, 2001
35
Histone modification-dependent recruitment of
proteins
Transcriptional activation
TAFII250
Bromodomain
36
Histone modification-dependent recruitment of
proteins
Heterochromatin assembly, Transcriptional
silencing
Transcriptional activation
HP1
TAFII250
Chromodomain
Bromodomain
Me
Ac
ARKSTGGK
...
...
H3
9
14
37
Histone methylation is important for defining
and maintaining epigenetic states
38
Identifying methyl-H3 binding proteins
- histone peptide pulldown assay
b
a
?
b
b
a
?
39
Site specific methylation of the H3 tail has
different functions
HP1
polycomb
BPTF
CD
CD
PhD
transcriptional competence
transcription repression
transcription repression
constitutive heterochromatin
euchromatin
facultative heterochromatin
40
Heterochromatin and euchromatin
constitutive heterochromatin
facultative heterochromatin
euchromatin
K9-methylated H3
K27-methylated H3
K4-methylated H3
HP1
polycomb
BPTF Yng2
41
Different dynamics of histone modifications
highly dynamic
HMT
more stable
histone
Me-histone
de-methylase
42
The search for histone demethylases
- LSD1 is a transcriptional co-repressor and its
repression function is - mediate through the amine-oxidase domain
Transcription ?
luciferase
5X Gal4 binding sites
Shi et al, Cell, 2004
43
The search for histone demethylases
- LSD1 is a histone H3-K4 demethylase
Shi et al, Cell, 2004
44
The search for histone demethylases
- LSD1 is a histone H3-K4 demethylase
Shi et al, Cell, 2004
45
The search for histone demethylases
Adapted from Tsukada and Zhang, Methods, 2006
46
Purifcation of histone demethylases
Release of radioactive formaldehyde
Adapted from Tsukada and Zhang, Methods, 2006
47
Identifying site of histone demethylation
- JHDM1A demethylates di-MeK36 on H3
Adapted from Tsukada and Zhang, Methods, 2006
48
Overexpression of JHDM1A results in loss of
K36Me-H3
Adapted from Tsukada and Zhang, Methods, 2006
49
Histone de-methylases are found for all these
sites
LSD1 JARID1a-d
JMJD2b
UTX JMJD3
Apart from LSD1, all other histone de-methylases
identified so far belong to the JmjC
domain-containing family of enzymes
50
Epienetics and diseases
diseases
- b-globin thalassemia
- leukemia
adapted from Nature 429, 2004
51
Paper assignments for Nov 2nd
Group 1 (Chanda - Fasih)
Kuzmichev et al, 2002, Genes Dev. 16
2893-2905 Histone methyltransferase activity
associated with a human multiprotein complex
containing the enhancer of Zeste protein
Group 2 (Fenton - How)
de Napoles et al, 2004, Dev. Cell 7
663-676 Polycomb group proteins Ring1A/B link
ubiquitylation of histone H2A to heritable gene
silencing and X inactivation
Group 3 (Karisch - Yan)
Wysocka et al, 2006, Nature 442 86-90 A PHD
finger of NURF couples histone H3 lysine 4
trimethylation with chromatin remodeling